Add this single word to make your Pandas Apply faster

We as data scientists have got laptops with quad-core, octa-core, turbo-boost. We work with servers with even more cores and computing power.
But do we really utilize the raw power we have at hand?
Sometimes we get limited by the limitation of tools at our disposal. And sometimes we are not willing to write all that extraneous code to save a couple of minutes. And end up realizing only later that time optimization would have helped in the long run.
So, can we do better?
Yes, Obviously.
Previously, I had written on how to make your apply function faster-using multiprocessing , but thanks to the swifter library, it is even more trivial now.
This post is about using the computing power we have at hand and applying it to Pandas DataFrames using Swifter.
Problem Statement
We have got a huge pandas data frame, and we want to apply a complex function to it which takes a lot of time.
For this post, I will generate some data with 25M rows and 4 columns.
Can use parallelization easily to get extra performance out of our code?
import pandas as pd
import numpy as np
pdf = pd.DataFrame(np.random.randint(0,100,size=(25000000, 4)),columns=list('abcd'))
The Data looks like:
 Data Sample
Data Sample
Parallelization using just a single change
 Relax and Parallelize !!!
Relax and Parallelize !!!
Let’s set up a simple experiment.
We will try to create a new column in our dataframe. We can do this simply by using apply-lambda in Pandas.
def func(a,b):
if a>50:
return True
elif b>75:
return True
else:
return False
pdf['e'] = pdf.apply(lambda x : func(x['a'],x['b']),axis=1)

The above code takes ~10 Minutes to run. And we are just doing a simple calculation on 2 columns here.
Can we do better and what would it take?
Yes, we can do better just by adding a “magic word” — Swifter.
But first, you need to install swifter, which is as simple as:
conda install -c conda-forge swifter
You can then just import and append swifter keyword before the apply to use it.
import swifter
pdf['e'] = pdf.**swifter**.apply(lambda x : func(x['a'],x['b']),axis=1)
So, Does this work?

Yes. It does. We get a 2x improvement in run time vs. just using the function as it is.
So what exactly is happening here?
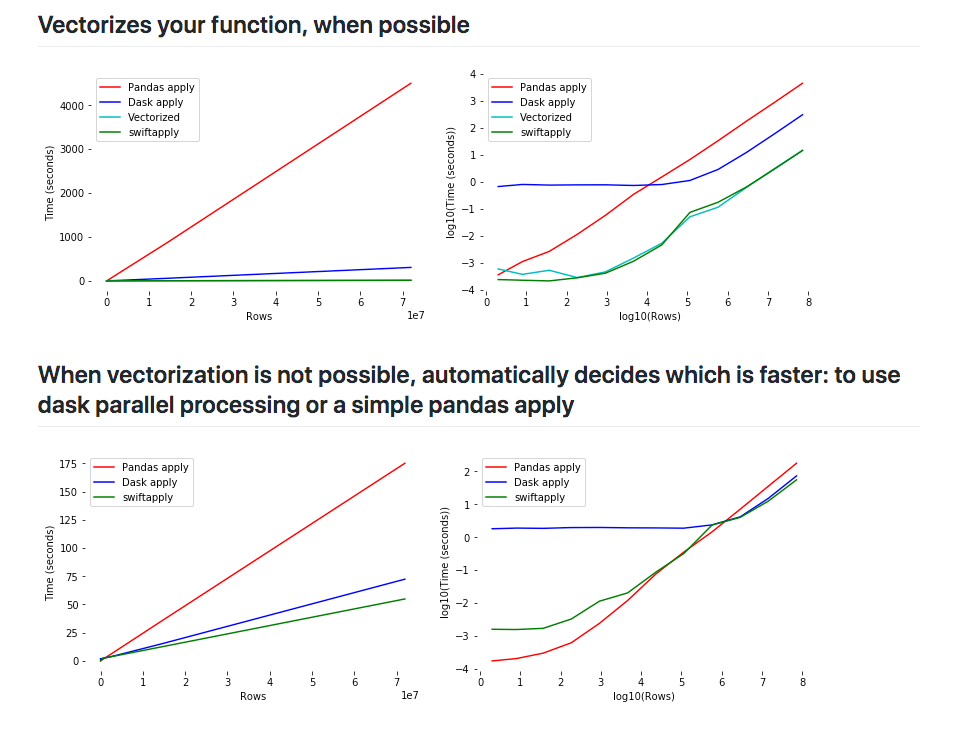 Source
: How increasing data size effects performances for Dask, Pandas and Swifter?
Source
: How increasing data size effects performances for Dask, Pandas and Swifter?
Swifter chooses the best way to implement the apply possible for your function by either vectorizing your function or using Dask in the backend to parallelize your function or by maybe using simple pandas apply if the dataset is small.
In this particular case, Swifter is using Dask to parallelize our apply functions with the default value of npartitions = cpu_count()*2.
For the MacBook I am working on the CPU Count is 6 and the hyperthreading is 2. Thus CPU Count is 12 and that makes npartitions=24.
We could also choose to set n_partitions ourselves. Though I have observed the default value works just fine in most cases sometimes you might be able to tune this as well to gain additional speedups.
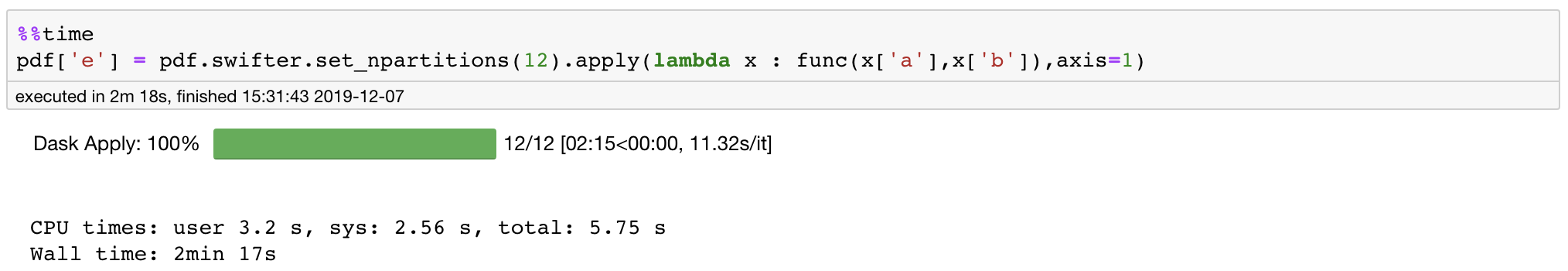
For example: Below I set n_partitions=12 and we get a 2x speedup again. Here reducing our number of partitions results in smaller run times as the data movement cost between the partitions is high.
Conclusion
Parallelization is not a silver bullet; it is buckshot.
Parallelization won’t solve all your problems, and you would still have to work on optimizing your functions, but it is a great tool to have in your arsenal.
Time never comes back, and sometimes we have a shortage of it. At these times we need parallelization to be at our disposal with a single word.
And that word is swifter .
Continue Learning
Also if you want to learn more about Python 3, I would like to call out an excellent course on Learn Intermediate level Python from the University of Michigan. Do check it out.
I am going to be writing more beginner-friendly posts in the future too. Follow me up at <strong>Medium</strong> or Subscribe to my <strong>blog</strong> .
Also, a small disclaimer — There might be some affiliate links in this post to relevant resources, as sharing knowledge is never a bad idea.
